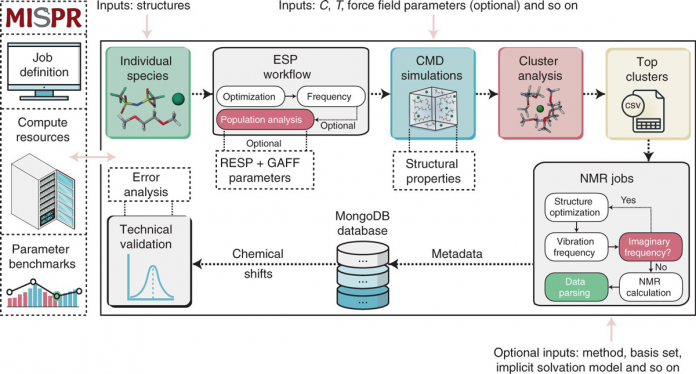
Stony Brook University researchers have found a way to computationally predict stable molecular species in liquid solutions. The new method has been detailed in a paper in Nature Computational Science. It introduces a fully automated high-throughput computational framework to predict stable species by computing their nuclear magnetic resonance chemical shifts.
Liquid solutions are essential aspects of both materials’ science applications such as battery development and biological applications such as drug discovery. An understanding of the structure and thermodynamic stability and transport of any chemical species in a solution is paramount, to optimize the performance of liquid applications. Scientists used NMR as a powerful technique to develop an atomistic view to predict stable species.
Scientists elucidated the complex solvation environment in multi-component liquid solutions can be a daunting task. They used advanced experimental and computational techniques. They combined density functional theory with classical molecular dynamics simulations to make our predictions via NMR.
Scientists developed and tested a computational framework that through simulations robustly and efficiently calculates, analyses, and stores NMR chemical shifts from a variety of molecules in liquid solutions. They say the framework addresses some of the most challenging aspects of computational NMR by eliminating intuition driven prediction of stable species in liquid solutions.
They add that data collected from this framework should provide fingerprints to guide future experimental investigations of liquid solutions. It has optimal properties for material and science applications.